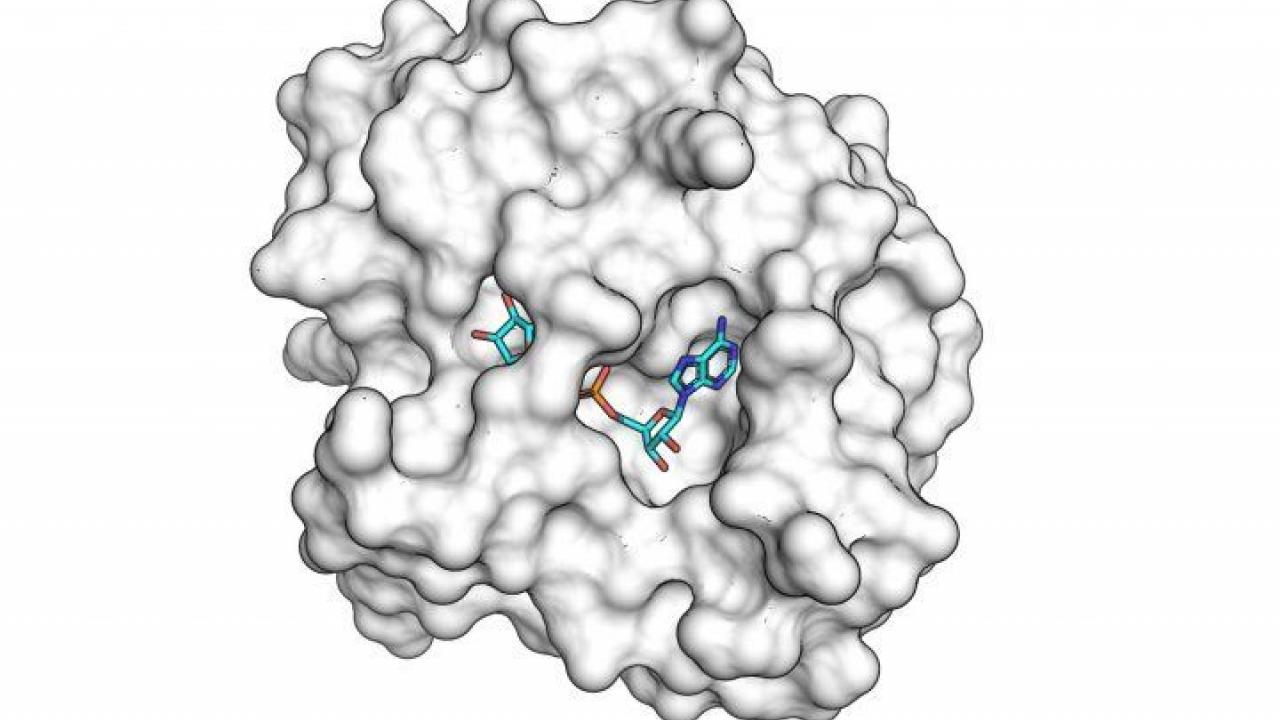
The SARS-CoV-2 macro domain protein bound to small molecule fragments that could be the basis of novel antiviral drugs.
Scientists have identified key chemical building blocks for an eventual antiviral drug against SARS-CoV-2, the virus that causes COVID-19, according to data newly released by members of the UC San Francisco Quantitative Bioscience Institute Coronavirus Research Group (QCRG), based on research conducted with breakneck speed in collaboration with Lawrence Berkeley National Laboratory (Berkeley Lab) and SLAC National Accelerator Laboratory (SLAC).
The newly identified compounds bind to an enzyme produced by the virus, called the “macro domain,” which is known to be crucial for the virus’s ability to replicate in human cells.
“The SARS-CoV-2 macro domain is not as well understood as the virus’s main protease or the spike proteins that other efforts are going after,” said James Fraser, PhD, an associate professor in the Department of Bioengineering and Therapeutic Sciences, based in the UCSF schools of Pharmacy and Medicine, who led the research effort as part of the UCSF QCRG Structural Biology Consortium. “We wanted to look where others aren’t looking as heavily and have succeeded in identifying some promising candidates to build drugs that could halt the virus’s ability to replicate and spread in the human body.”
Further work is needed to weld these potential active ingredients into a workable drug candidate, but the research represents a promising new frontier in the battle against the virus, the researchers say, particularly given uncertainties about the timing or ultimate efficacy of an eventual vaccine.
The authors are writing up a formal manuscript describing the results for submission to a peer-reviewed academic journal, but also published their data directly online on July 1, 2020, to accelerate global efforts to fight the coronavirus pandemic. The open dataset was published in coordination with the Diamond Light Source X-Chem research group and Ivan Ahel’s lab at based at Oxford University, which simultaneously published their own parallel, complementary dataset of macrodomain-bound fragments.
“We’re delighted to openly share this data in coordination with the Diamond Light Source team, whose publication of a structure and fragment analysis of the SARS-CoV-2 main protease earlier this year was a huge step forward in galvanizing the computational chemistry community to develop additional antiviral targets,” Fraser said.
Novel Line of Attack on Coronavirus Biology
The study focused on an aspect of coronavirus biology that has only recently begun to be studied by researchers, including Alan Ashworth, PhD, FRS, president of the UCSF Helen Diller Family Comprehensive Cancer Center and a world leader in research on PARPs, enzymes that play a major role in DNA repair and have been targeted by drugs to treat certain cancers.
“PARPs can also reduce the effects of viral infection by tagging proteins with a molecule called ADP-ribose. As a countermeasure, some viruses – including the SARS family – have evolved macro domains that can remove this tag, re-exposing cells to infection,” Ashworth explained. “We are working to produce a drug to block the macro domain’s ability to remove these protective tags with the aim of reducing the impact of SARS-CoV-2 infection.”