The Molecular Oncology Initiative has launched a new version of the UCSF cBioPortal that links de-identified UCSF500 cancer sequencing panel data, to de-identified clinical data extracted from APEX and available through the Clinical Data Warehouse (CDW). Tests results from patients sequenced using Foundation Medicine are also available and have been linked to CDW.
The portal currently includes vital status, race and ethnicity derived from de-identified CDW with plans to include additional clinical data elements including treatment information and major cancer progression events. The UCSF cBioPortal shares the same patient identifiers as de-identified CDW, allowing users to define molecular cohorts of interest within cBioPortal and then direct querying of de-identified CDW tables for specific clinical data elements of interest.
What is cBioPortal?
Originally developed by Memorial Sloan Kettering Cancer Center, cBioPortal is an open-source platform that allows for visualization, exploration, and analysis of cancer genomic datasets. It has become a widely adopted, user-friendly tool for researchers throughout the world.
- User-defined cohorts
- Cohort summaries
- Patient/sample level discovery
- Supports genomic and clinical data
- Annotations from UniProt, OncoKb and others
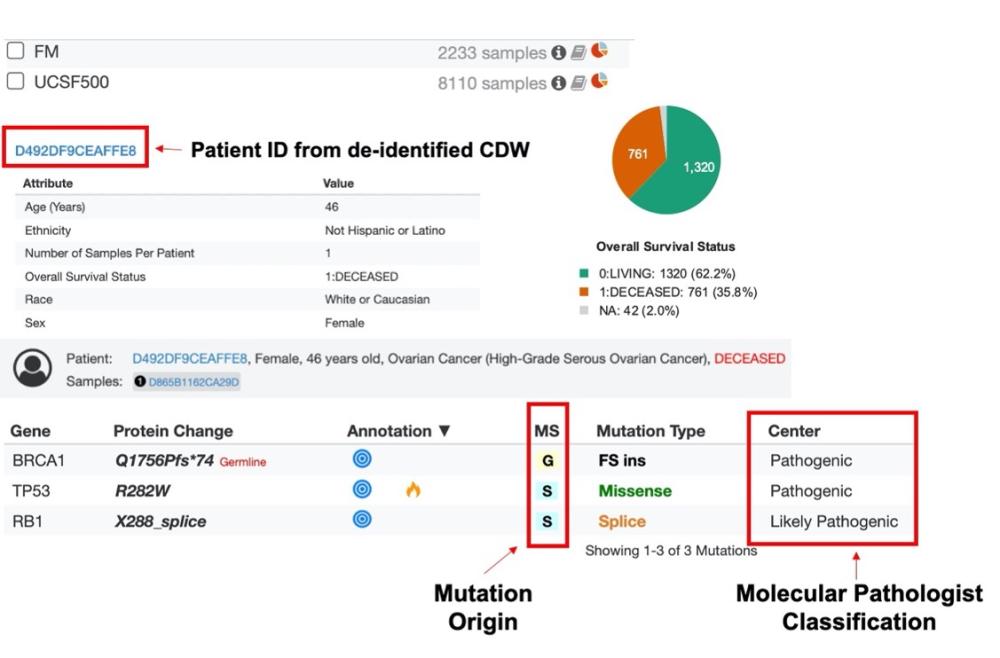
The newly developed and released pipeline that links to de-identified CDW and includes Foundation Medicine data in addition to UCSF500 data also provides significant improvements in the UCSF500 dataset: refinements in variant filtering to show clinically-relevant variants, an improved algorithm for inclusion of reported structural variants in the portal, cleaner variant origin annotations and the most recent assigned classification of a specific variant by our own expert molecular pathologists from the Clinical Cancer Genomics Laboratory.
How to get access
UCSF staff, faculty and students can access the UCSF cBioPortal. To obtain access, (1) fill-out and submit the Terms of Use agreement, (2) request a de-identified CDW account and (3) complete associated training modules for de-identified CDW (‘CyberSecurity Fundamentals’, ‘HIPAA101’, ‘Your Responsibilities when Working with Clinical Data’). Registered users can access the UCSF cBioPortal through MyAccess (cBioPortal – UCSF).
More questions about the UCSF cBioPortal, the data in the portal, how to get access or just feedback? Email [email protected]